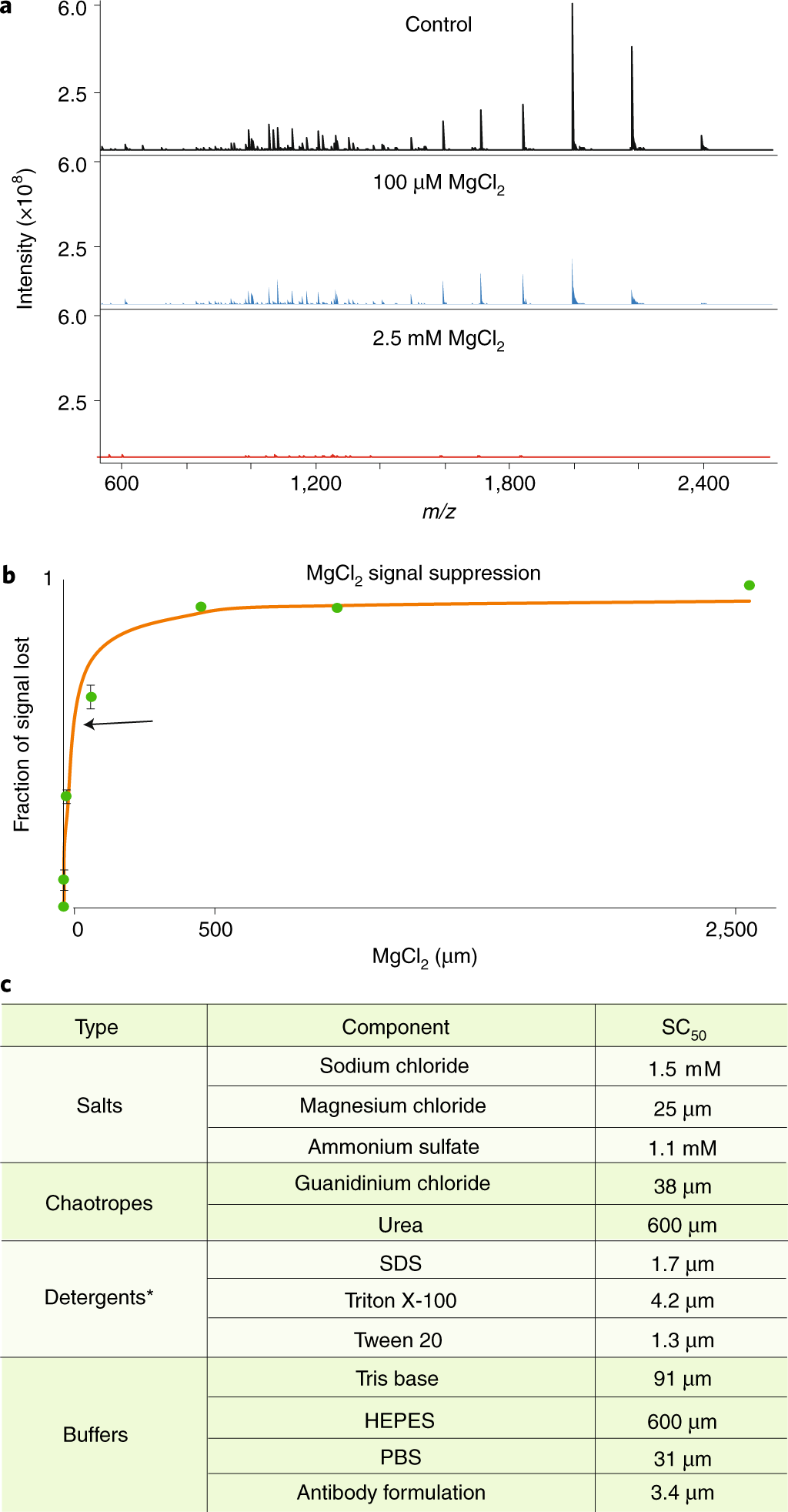
Best practices and benchmarks for intact protein analysis for top-down mass spectrometry | Nature Methods
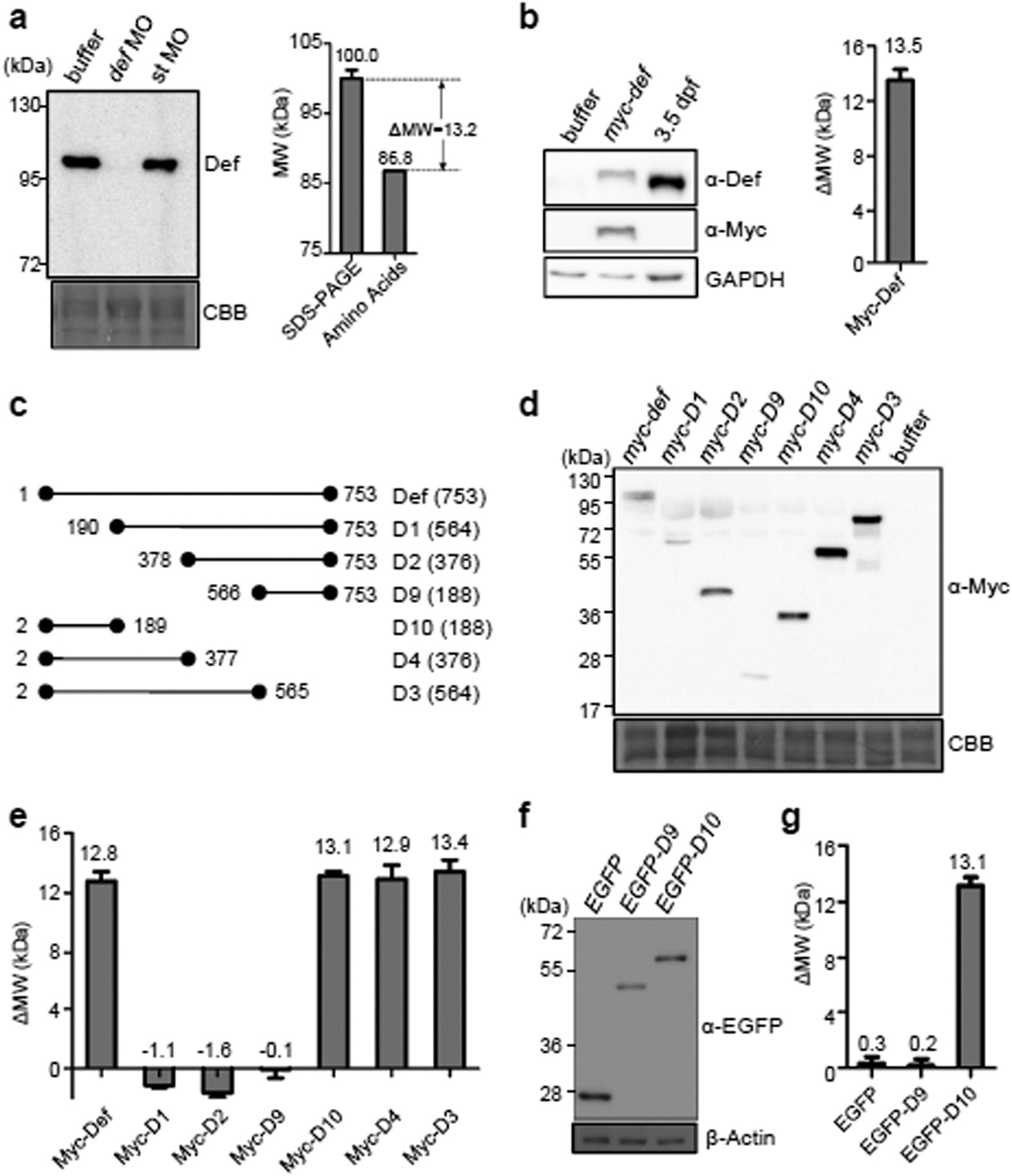
An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide | Scientific Reports
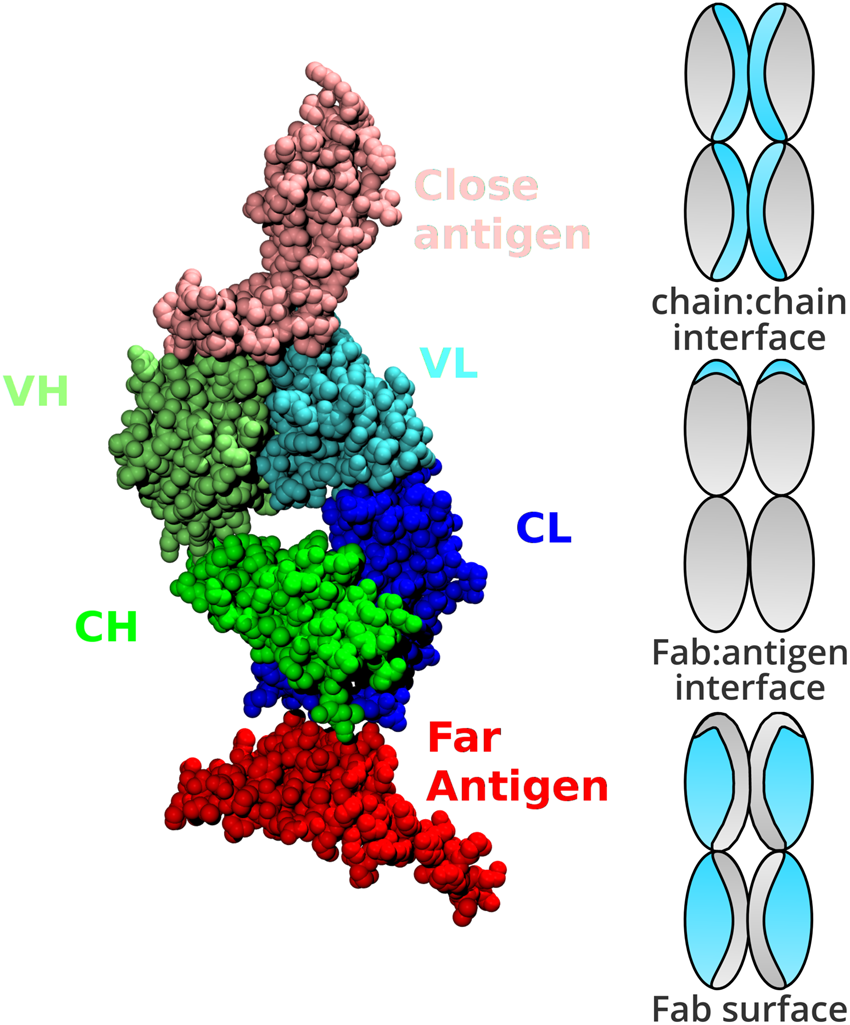
Web-based display of protein surface and pH-dependent properties for assessing the developability of biotherapeutics | Scientific Reports

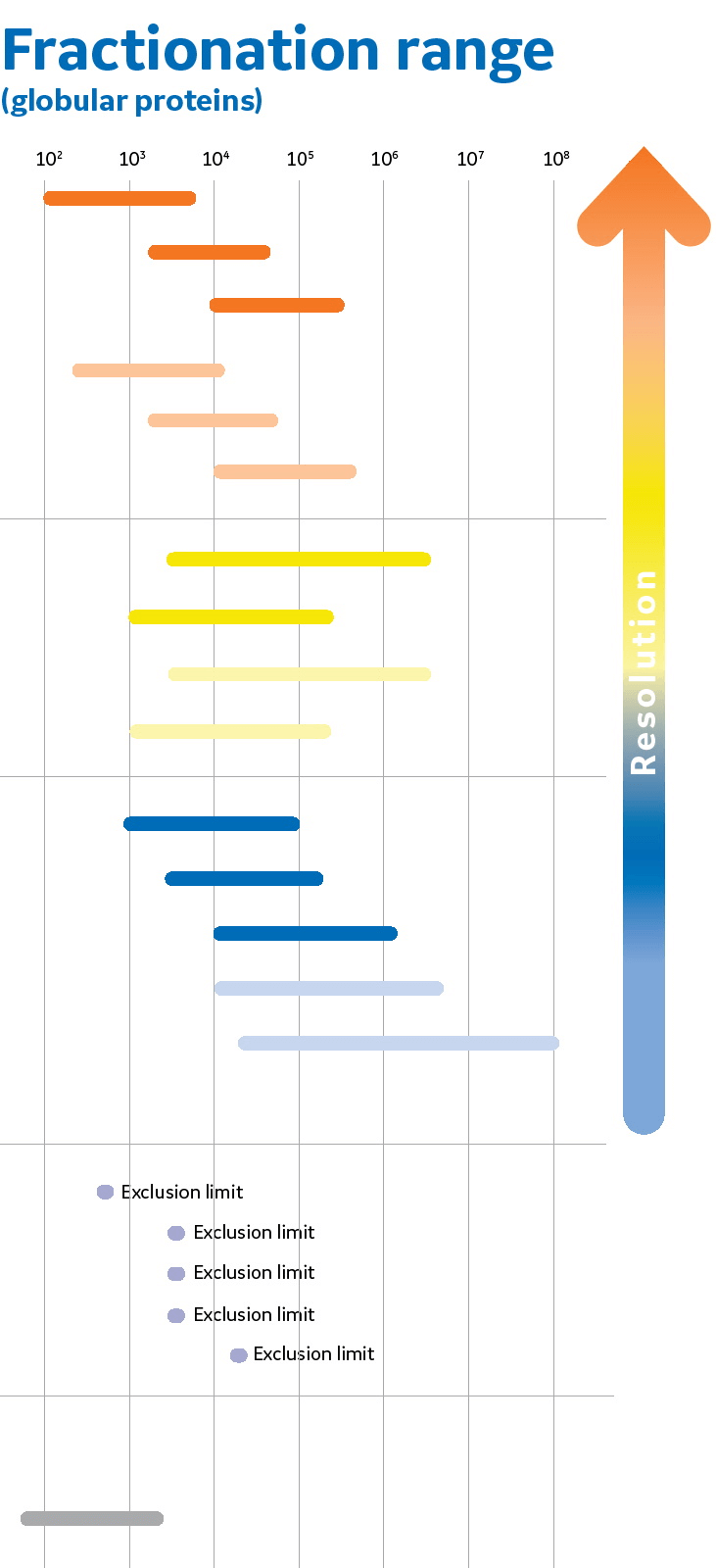
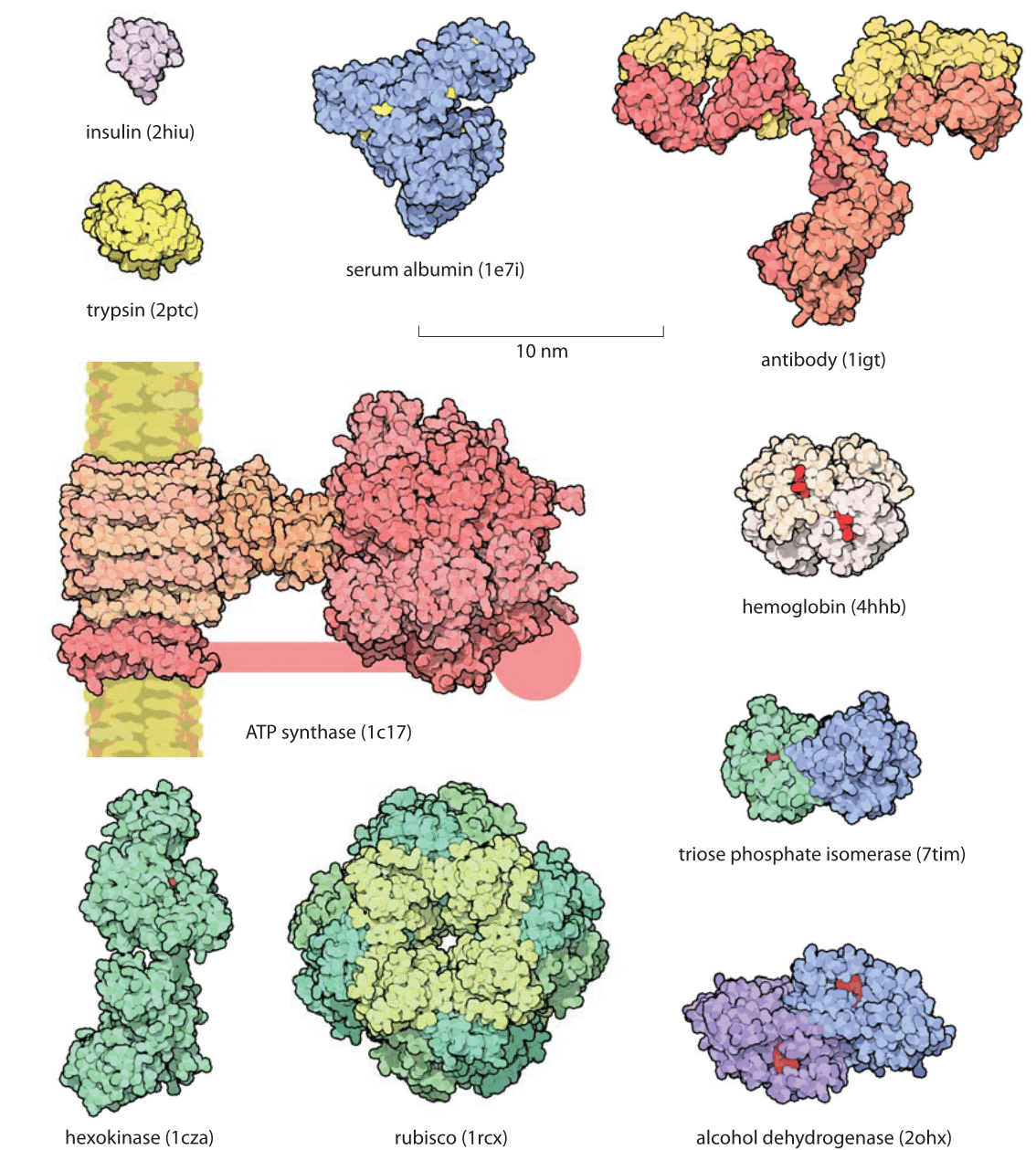
PDF) Size and Shape of Protein Molecules at the Nanometer Level Determined by Sedimentation, Gel Filtration, and Electron Microscopy

Why Does the Molecular Weight of My Protein Differ From the Theoretically Expected Weight? | Technology Networks
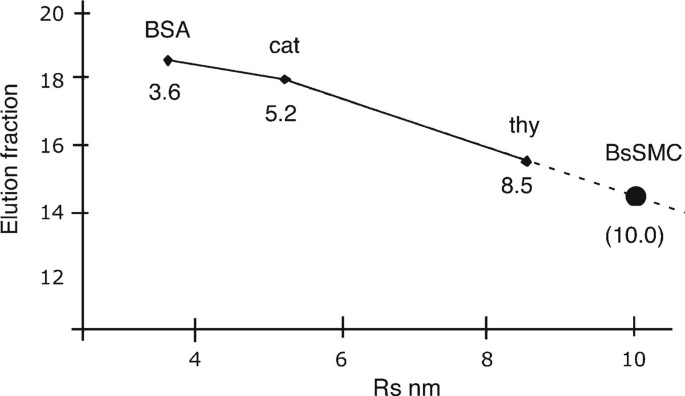
Size and Shape of Protein Molecules at the Nanometer Level Determined by Sedimentation, Gel Filtration, and Electron Microscopy | Biological Procedures Online | Full Text